Hunting for Viruses in French Guiana
I don’t know who that is, but it sure ain’t me.
The International Division of the Pasteur Institute has generously awarded me a grant to carry out a 4-month research internship in its branch located in French Guiana. The goal of the project is to determine the viral biodiversity of multiple species of rodents living in different habitats in the region.
Where I’m going
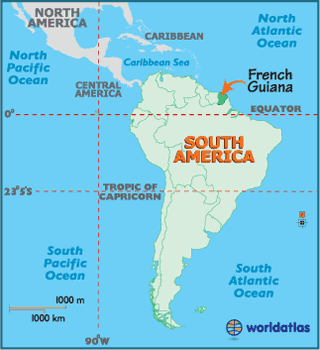
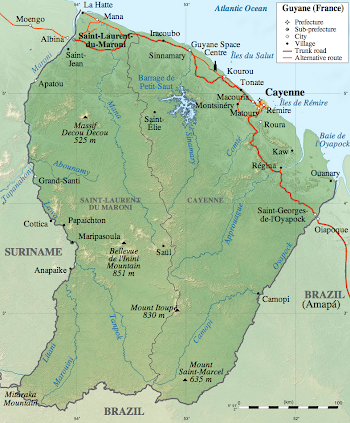
Although French Guiana borders Brazil and Suriname, it is not an independent country, but an overseas region of France. Therefore, it is part of the European Union, the official currency is the euro, and the license plates look European. The population is around 250,000 people, and half of them live in the capital, Cayenne. According to Wikipedia, there is one road that crosses the country parallel to the coast, and the rest is Amazonian jungle.
The Pasteur Institute in French Guiana was created in 1940 and it employs around 100 scientists, distributed across 11 different labs. I will be working in the Virus-Host Interactions lab.
What I’m doing
Whenever we urbanize or adapt a terrain for agriculture, we run the risk of disturbing the habitat of the species that live in it. This increases the chances that species that don’t usually interact will come into contact with each other; and raises the possibility that these interactions will help pathogens cross the species barrier, thereby expanding their range. Many different species of rodents live in close proximity to humans, but we don’t know much about the viruses they harbor. The little we know is that three rodent species are the likely reservoirs of some nasty viruses that killed several patients in French Guiana last year.
Metagenomics is a cool bioinformatics technique that can be used to determine all the genetic information that exists in a biological sample. This means that we can go capture rodents all over the country, take a few blood samples, process them and isolate what is likely to viral genetic material. Then, we sequence it and somebody gives me a hard drive full of As, Cs, Gs, and Ts—which is when I come in, because I’m not excited about getting bitten by virus-laden jungle rats.
My job is to determine what sequences are likely to belong to previously known viruses, to identify which ones may correspond to novel viruses (typically 50% of all the generated sequences are completely unknown!), and which ones may be potential human pathogens, and also, to compare the viral families in each sample and see if they correlate better by species or by habitat. Cool stuff.